Bryan T. GrenfellPopulation BiologistBasic Sciences
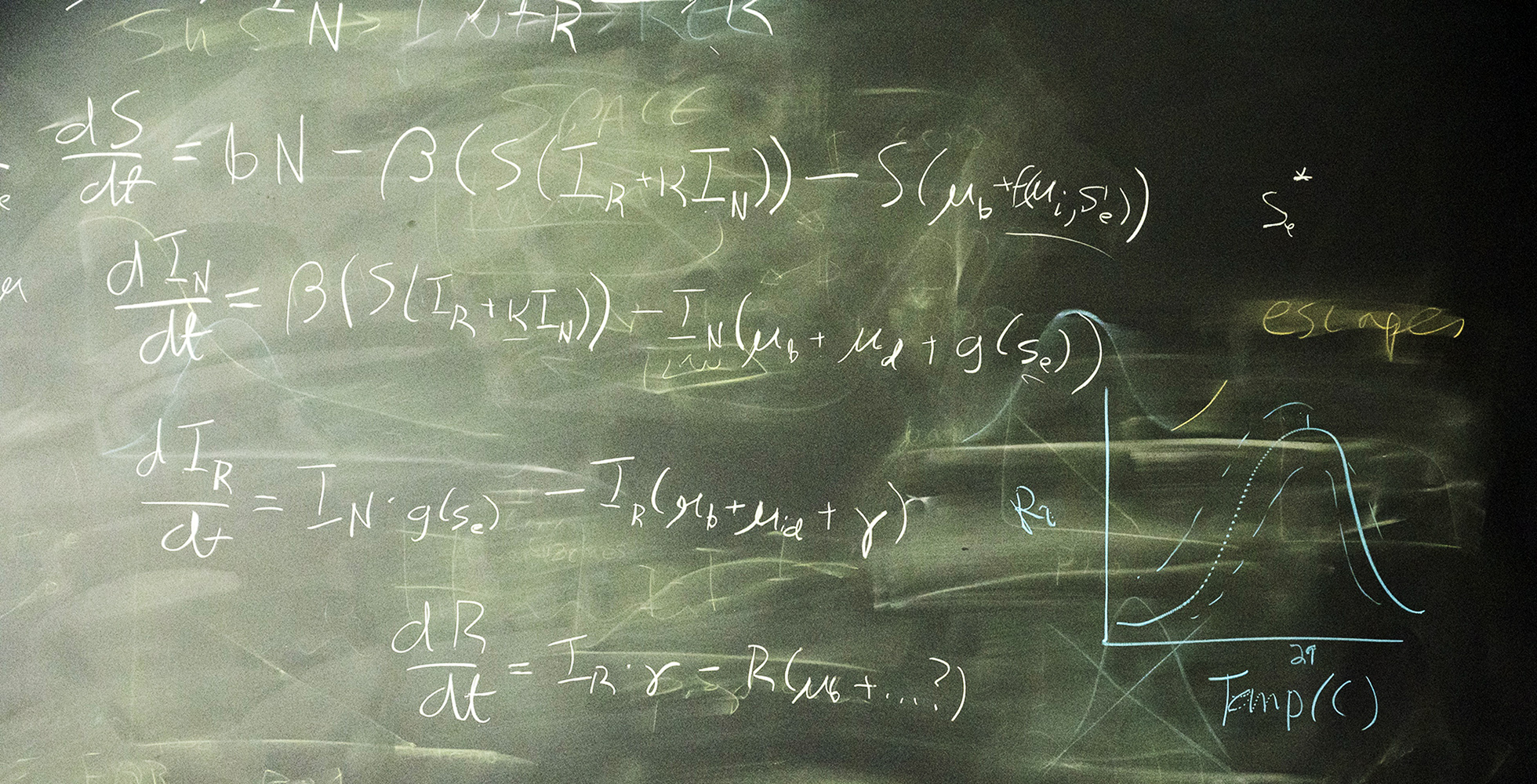
How can we understand infectious disease dynamics using mathematics?
Development of an Innovative Methodology for Integrative Analysis of Pathogen Evolution and Epidemics
Bryan T. Grenfell/ Population Biologist
Bryan T. Grenfell proposed “phylodynamics,” a methodology that predicts infectious disease dynamics of RNA viruses by considering viral evolution, and thus contributed to the development of the research field that integrates immune dynamics, epidemiology, and evolutionary biology. By virtue of these achievements, he has been instrumental in understanding infection mechanisms and proposing effective infectious disease control policies.
KEYWORDS
- #mathematical model
- #pathogen evolution
- #prediction of infectious disease dynamics
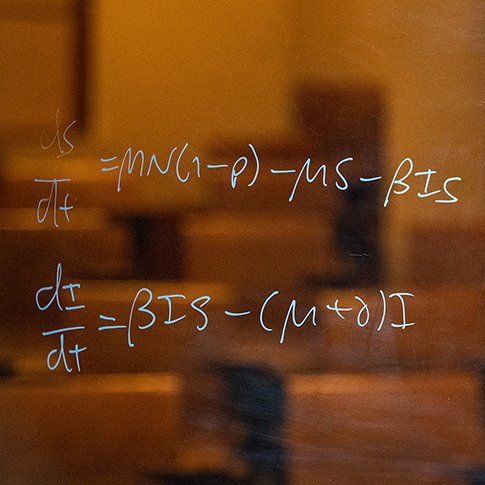


Introduction
Commemorative Lecture
Epidemiological and Evolutionary Dynamics of Pathogens in Time and Space
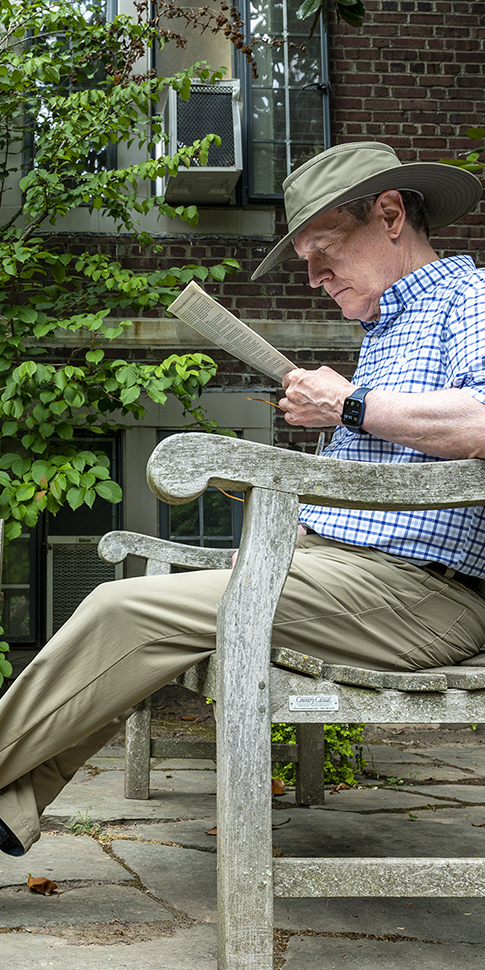
I have spent my career exploring the population dynamics and evolution of infectious diseases. I begin this lecture by introducing epidemics and the rich historical data sets which illuminate their dynamics. I focus on measles as a clear exemplar of the epidemic dynamics of acute immunizing infections. I then describe my research, under three main themes. First, I and collaborators explored the non-linear temporal dynamics of childhood infections, notably measles. We used simple mathematical models and time series analysis of epidemic data to explain oscillatory, sometimes chaotic epidemic sequences and the impact of seasonal and demographic drivers and vaccination on these patterns. Second, we analyzed the spatio-temporal dynamics of regional epidemics and the determinants of local epidemic persistence. Third, we explored the evolutionary dynamics of partly-immunizing pathogens such as influenza and SARS-CoV-2. I coined the term “phylodynamics” for the interactions between viral evolution and epidemiology that drives the “escape” of viral variants from prevailing immunity; phylodynamic ideas have since been applied to a wide variety of problems in pathogen evolution. I conclude by drawing broader lessons from my career, notably the power and pleasure of interdisciplinary collaboration.
Biology is often extremely complex but, sometimes, simple models can explain some of the complexity.

Bryan T. Grenfell/ Population Biologist
Profile
Biography
- 1954Born in Swansea, U.K.
- 1981D.Phil., University of York
- 1981–1986Research Fellow, Department of Pure and Applied Biology, Imperial College London
- 1986–1990Lecturer, Department of Animal and Plant Sciences, University of Sheffield
- 1990–1998Lecturer, Department of Zoology, University of Cambridge
- 1998–2002Reader, Department of Zoology, University of Cambridge
- 2002–2004Professor of Population Biology, Department of Zoology, University of Cambridge
- 2004–2009Alumni Professor of Biology, Pennsylvania State University
- 2009–Kathryn Briger and Sarah Fenton Professor of Ecology and Evolutionary Biology and Public Affairs, Department of Ecology and Evolutionary Biology and Princeton School of Public and International Affairs, Princeton University
- 2014–2021Member of Governing Board, Wellcome Trust
Selected Awards and Honors
- 1991T.H. Huxley Medal of Imperial College, London
- 1995Scientific Medal of the Zoological Society of London
- MembershipsAmerican Academy of Arts and Sciences, American Association for the Advancement of Science, Royal Society
Profile is at the time of the award.
- Advanced Technology
- Basic Sciences
- Arts and Philosophy

